This article is more than 1 year old
Y'know how you might look at someone and can't help but wonder if they have a genetic disorder? We've taught AI to do the same
More suggestive assistant than robo-doc, boffins say
Artificial intelligence can potentially identify someone's genetic disorders by inspecting a picture of their face, according to a paper published in Nature Medicine this week.
The tech relies on the fact some genetic conditions impact not just a person’s health, mental function, and behaviour, but sometimes are accompanied with distinct facial characteristics. For example, people with Down Syndrome are more likely to have angled eyes, a flatter nose and head, or abnormally shaped teeth. Other disorders like Noonan Syndrome are distinguished by having a wide forehead, a large gap between the eyes, or a small jaw. You get the idea.
An international group of researchers, led by US-based FDNA, turned to machine-learning software to study genetic mutations, and believe that machines can help doctors diagnose patients with genetic disorders using their headshots.
The team used 17,106 faces to train a convolutional neural network (CNN), commonly used in computer vision tasks, to screen for 216 genetic syndromes. The images were obtained from two sources: publicly available medical reference libraries, and snaps submitted by users of a smartphone app called Face2Gene, developed by FDNA.
Given an image, the system, dubbed DeepGestalt, studies a person's face to make a note of the size and shape of their eyes, nose, and mouth. Next, the face is split into regions, and each piece is fed into the CNN. The pixels in each region of the face are represented as vectors and mapped to a set of features that are commonly associated with the genetic disorders learned by the neural network during its training process.
DeepGestalt then assigns a score per syndrome for each region, and collects these results to compile a list of its top 10 genetic disorder guesses from that submitted face.
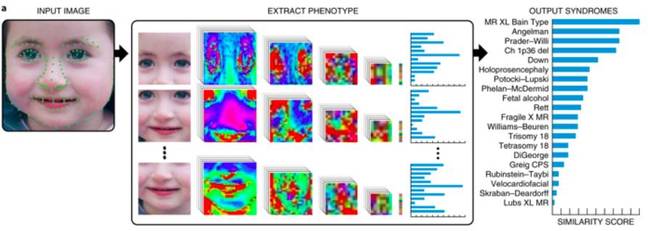
An example of how DeepGestalt works. First, the input image is analysed using landmarks and sectioned into different regions before the system spits out its top 10 predictions. Image credit: Nature and Gurovich et al.
The first answer is the genetic disorder DeepGestalt believes the patient is most likely affected by, all the way down to its tenth answer, which is the tenth most likely disorder.
When it was tested on two independent datasets, the system accurately guessed the correct genetic disorder among its top 10 suggestions around 90 per cent of the time. At first glance, the results seem promising. The paper also mentions DeepGestalt “outperformed clinicians in three initial experiments, two with the goal of distinguishing subjects with a target syndrome from other syndromes, and one of separating different genetic subtypes in Noonan Syndrome.”
There's always a but
A closer look, though, reveals that the lofty claims involve training and testing the system on limited datasets – in other words, if you stray outside the software's comfort zone, and show it unfamiliar faces, it probably won't perform that well. The authors admit previous similar studies “have used small-scale data for training, typically up to 200 images, which are small for deep-learning models.” Although they use a total of more than 17,000 training images, when spread across 216 genetic syndromes, the training dataset for each one ends up being pretty small.
For example, the model that examined Noonan Syndrome was only trained on 278 images. The datasets DeepGestalt were tested against were similarly small. One only contained 502 patient images, and the other 392.
It'll be difficult to reproduce and verify the results because the Face2Gen-sourced photographs used to train and test the neural network are protected, and cannot be shared with other research efforts. On top of that, beyond the public references, it's tricky to get hold of actual patients' faces ethically, making datasets scarce, if you wanted to create your own model from scratch. The public datasets used to teach DeepGestalt are listed in table six of this PDF.
Yaron Gurovich, lead author of the study and a researcher at FNDA, told The Register: "Every AI system is eventually evaluated by some benchmark. We evaluate on two different benchmarks, and we make one available to the public. So researchers are welcome to take a closer look on this benchmark to see the detailed performance. The training data is hard to obtain, as this training data consists of real patients from diverse locations worldwide."
What’s worse is that the researchers don’t really know how DeepGestalt arrives at its answer. “DeepGestalt, like many artificial intelligence systems, cannot explicitly explain its predictions and provides no information about which facial features drove the classification,” the team's paper stated.
Both training and testing datasets are largely made up of faces from Caucasian backgrounds, and the algorithm's performance may differ for people of other races. "It is possible to extend this system to a wide range of ethnic backgrounds. We are extending our system to be able to support this functionality," Gurovich said.
At the moment, some genetic disorders are diagnosed by doctors studying a patient's physical health and characteristics. The researchers argue that this method is dependent on a clinician’s previous experience, and having an automated approach based on facial analysis could offer “better syndrome prioritization and diagnosis." To put it another way, if a doc is going by looks and health alone, a computer system may be more consistent in identifying disorders.
Gurovich explained to El Reg, however, that the software shouldn't really be used to make a final decision, and should be used more as a reference for doctors. "We trust the experts to perform the diagnostic process while the goal of our system is to provide them with richer and wider tools and information."
DeepGestalt, don't forget, cannot outright predict if a patient has a disorder or not. It instead outputs a list of which genetic syndromes it believes a person may have even if they aren't affected with any. "The task of predicting if a patient has a disorder or not is not in the scope of our work," said Gurovich.
This type of research is still very new and since there are no standardized tests, it’s impossible to check the performance and accuracy of the system against other models. The code is also proprietary, although some of the resources used to train the system are available through the Face2Gene database.
"Our results open the door for an extended use of the phenotyping to improve the diagnostic yield of genomics. Our Face2Gene system is now used within bioinformatics platforms," Gurovich told us. ®

